Celina
Lulu Shang & Peijun Wu
2024-03-19
Last updated: 2024-03-21
Checks: 2 0
Knit directory: Celina/
This reproducible R Markdown analysis was created with workflowr (version 1.7.1). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Untracked files:
Untracked: .Rprofile
Untracked: .gitattributes
Untracked: .gitignore
Untracked: Celina.Rproj
Untracked: README.md
Untracked: _workflowr.yml
Untracked: analysis/
Untracked: code/
Untracked: data/
Untracked: output/
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with
wflow_publish() to start tracking its development.
Welcome to Celina!
Overview
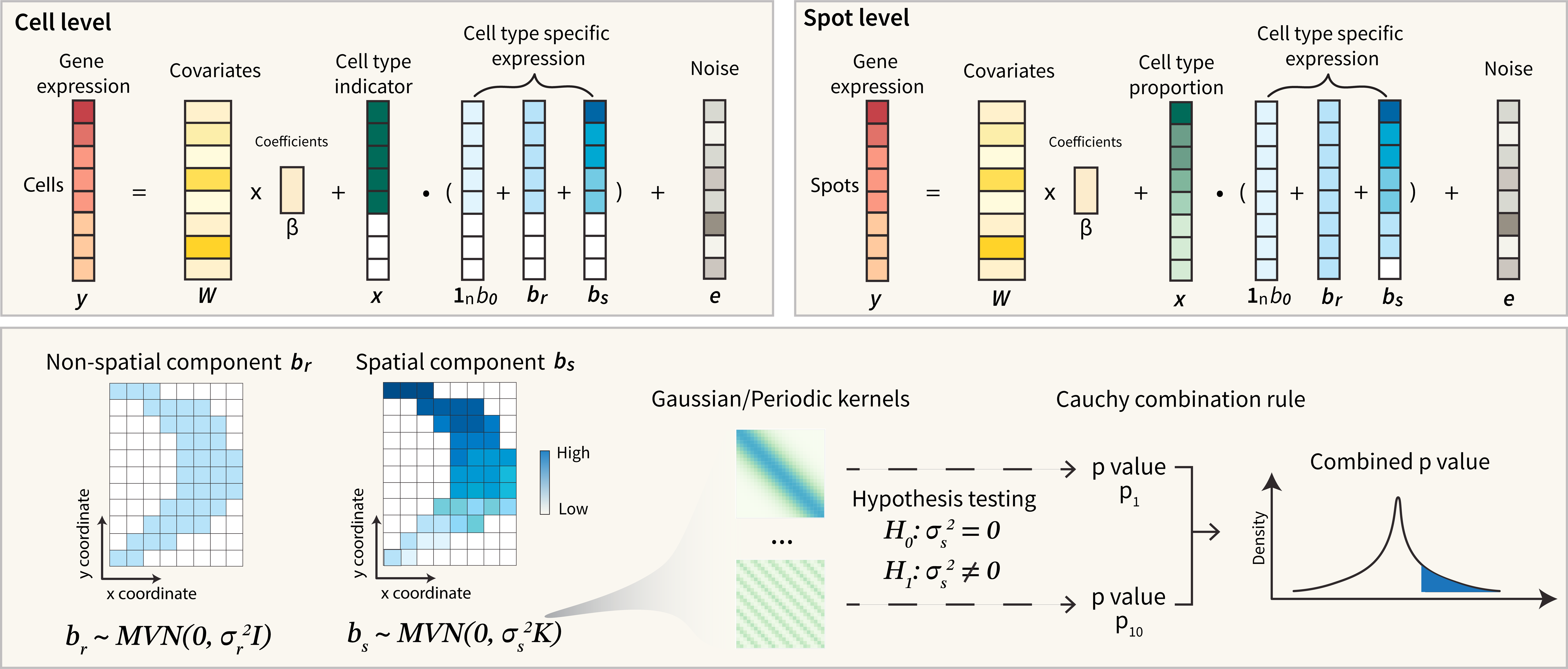
Celina a statistical method for systematically detecting cell type-specific spatially variable genes (ct-SVGs). Celina utilizes a spatially varying coefficient model to accurately capture each gene’s spatial expression pattern in relation to the distribution of cell types across tissue locations, ensuring effective type I error control and high statistical power.